The molecular basis of plant growth & development
Molecular Genetics & Natural Genetic Variation
A central feature of plant development is the post-embryonic formation of organs in a reiterative fashion. This build-up of the body continues until the plant dies and can last hundreds of years in long-lived species, like certain trees. Thus, control of plant growth and elaboration of plant form are central to plant development. Our research revolves around the genetic control of these processes, and the underlying molecular mechanisms. A major focus of our lab is vascular differentiation and its relation to plant hormones.
The importance of the vascular system for plant development cannot be overstated. Its evolution enabled plants to effectively colonize land and thus had a long-lasting impact that shaped earth history and the extant biosphere. Vascular tissues allowed body plan expansion, because they permit long distance separation of the location of water and nutrient acquisition from the location of photosynthesis. At the heart of the vasculature, xylem transports water and inorganic nutrients absorbed by the root system to aboveground organs, whereas phloem distributes photosynthetic and other organic metabolites throughout the plant. Xylem and phloem are continuously formed during organ formation from stem cell niches in plant meristems, the growth apices of plant body axes. It is in there where we investigate the molecular details and evolutionary aspects of pathways that are required for proper vascular tissue formation. Our favorite experimental organisms are the dicotyledon model Arabidopsis thaliana, and the monocotyledon model, Brachypodium distachyon.
Recent Publications
- BAM1/2 receptor kinase signaling drives CLE peptide-mediated formative cell divisions in Arabidopsis roots
Crook, AD; Willoughby, AC; Hazak, O; Okuda, S; VanDerMolen, KR; Soyars, CL; Cattaneo, P; Clark, NM ; Sozzani, R; Hothorn, M; Hardtke, CS; Nimchuk, ZL
Proceedings Of The National Academy Of Sciences Of The United States Of America 10.1073/pnas.2018565117 DEC 22 2020 - Arabidopsis Flippases Cooperate with ARF GTPase Exchange Factors to Regulate the Trafficking and Polarity of PIN Auxin Transporters
Zhang, Xixi; Adamowski, Maciek; Marhava, Petra; Tan, Shutang; Zhang, Yuzhou; et al.
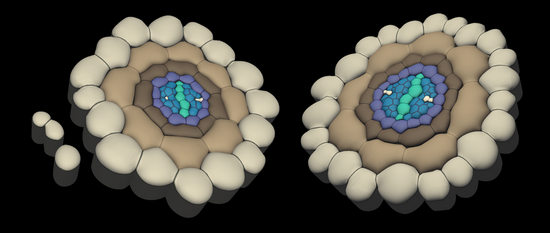
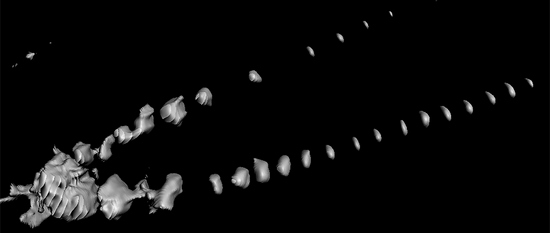
Plant Cell 10.1105/tpc.19.00869 MAY 2020 - Local auxin competition explains fragmented differentiation patterns
Moret, Bernard; Marhava, Petra; Fandino, Ana Cecilia Aliaga; Hardtke, Christian S.; ten Tusscher, Kirsten H. W.
Nature Communications 10.1038/s41467-020-16803-7 Published: JUN 11 2020 - The transcription and export complex THO/TREX contributes to transcription termination in plants
Khan, Ghazanfar Abbas; Deforges, Jules; Reis, Rodrigo S.; Hsieh, Yi-Fang; Montpetit, Jonatan; Antosz, Wojciech; Santuari, Luca; Hardtke, Christian S; Grasser, Klaus D; Poirier, Yves
Plos Genetics DOI: 10.1371/journal.pgen.1008732 Published: APR 2020 - Local and Systemic Effects of Brassinosteroid Perception in Developing Phloem
Graeff, Moritz; Rana, Surbhi; Marhava, Petra; Moret, Bernard; Hardtke, Christian S.
Current Biology 30(9) DOI: 10.1016/j.cub.2020.02.029 MAY 2020 - Plant Biology: Brassinosteroids and the Intracellular Auxin Shuttle
Rana, Surbhi; Hardtke, Christian S.
Current Biology 30(9) DOI: 10.1016/j.cub.2020.02.073 MAY 2020 - Peptide Signaling Pathways in Vascular Differentiation
Fukuda, Hiroo; Hardtke, Christian S.
Plant Physiology 182 (4)1636-1644 DOI: 10.1104/pp.19.01259 APR 2020 - Plasma Membrane Domain Patterning and Self-Reinforcing Polarity in Arabidopsis
Marhava, Petra; Fandino, Ana Cecilia Aliaga; Koh, Samuel W. H.; Jelinkova, A; Kolb, M; Janacek, DP; Breda, AS; Cattaneo, P; Hammes, UZ; Petrasek, J; Hardtke, CS
DEVELOPMENTAL CELL, https://doi.org/10.1016/j.devcel.2019.11.015, JAN 2020 - Conditional effects of the epigenetic regulator JUMONJI 14 in Arabidopsis root growth
Cattaneo, Pietro; Graeff, Moritz; Marhava, Petra; Hardtke, Christian S.
Development, 10.1242/dev.183905, DEC 2019 - A Cellular Insulator against CLE45 Peptide Signaling
Breda, AS; Hazak, O; Schultz, P; Anne, P; Graeff, M; Simon, R; Hardtke, CS
Current Biology, 29 (15):2501-+; 10.1016/j.cub.2019.06.037 AUG 5 2019 - Broad spectrum developmental role of Brachypodium AUX1
van der Schuren, Alja; Voiniciuc, Catalin; Bragg, Jennifer et al.
NEW PHYTOLOGIST, https://doi.org/10.1111/nph.15332, Jun 2018 - A molecular rheostat adjusts auxin flux to promote root protophloem differentiation
Marhava, P; Bassukas, AEL; Zourelidou, M; Kolb, M; Moret, B; Fastner, A; Schulze, WX; Cattaneo, P; Hammes, UZ; Schwechheimer, C; Hardtke, CS
NATURE, 10.1038/s41586-018-0186-z, JUN 14 2018 - CLERK is a novel receptor kinase required for sensing of root-active CLE peptides in Arabidopsis
Anne, Pauline; Amiguet-Vercher, Amelia; Brandt, Benjamin; Kalmbach, Lothar; Geldner, Niko; Hothorn, Michael; Hardtke, Christian S
DEVELOPMENT Volume: 145 Issue: 10 https://doi.org/10.1242/dev.162354 MAY 2018 - Phloem function and development — biophysics meets genetics
Pauline Anne and Christian S. Hardtke
Current Opinion in Plant Biology, Vol. 43: pp. 22-28. https://doi.org/10.1016/j.pbi.2017.12.005 (2018) - …

Prof. Dr. Christian Hardtke
University of Lausanne
Department of Plant Molecular Biology
1015 Lausanne
Tel: +41 (0)21 692 42 51
Research topics
- Natural genetic variation
- Root system growth & architecture
- Secondary growth
- Plant hormone pathways
Interdisciplinary
- Process modeling of root growth (in collaboration with Prof. Richard Smith, Bern; Dr. Ioannis Xenarios, SIB)
- Bioinformatics approaches (collaboration with Dr. Ioannis Xenarios, SIB)