Plant adaptation and speciation in the face of gene flow
Population Genomics
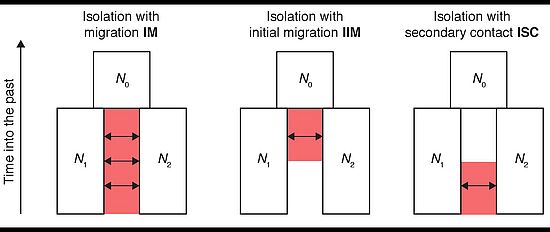
Speciation and adaptive divergence often include periods of gene flow. Whether this gene flow occurs prior, during, or after the evolution of reproductive isolation, and what consequences gene flow has on the architecture of traits under divergent selection are fundamental questions that remain largely unanswered.
While we now have unprecedented amounts of genome scale data, there is a lack of statistical approaches to make robust inference from these data. To fill this gap, we combine mathematical modelling and population genetic theory to devise statistical procedures for the joint inference of demography and selection. Applying these approaches to genome-wide sequence and recombination data, we ask how selection modifies gene flow in speciation, what genes act as barriers to gene flow, and how deleterious mutations accumulate in populations as a function of demographic history. We currently use Mimulus and wild tomatoes (Solanum section Lycopersicon) as our main study systems.
Research topics
- Inferring the timing and strength of gene flow and selection
- Understanding the genetic basis of adaptation and speciation
- Quantifying the impact of demography on natural selection
Recent Publications
- ORCID, https://orcid.org/0000-0001-5217-1926
- Comparative evaluation of the MAPlex, Precision ID Ancestry Panel, and VISAGE Basic Tool for biogeographical ancestry inference
P Resutik, S Aeschbacher, M Krützen, A Kratzer, C Haas, C Phillips, ...
Forensic Science International: Genetics 64, 102850 - Background selection and biased gene conversion affect more than 95% of the human genome and bias demographic inferences.
Poyet F, Aeschbacher S, Thiéry A, Excoffier L (2018).
eLife 2018;7:e36317. DOI: 10.7554/eLife.36317 - Population-genomic inference of the strength and timing of selection against gene flow.
Aeschbacher S, Selby JP, Willis JH, Coop G (2017).
Proceedings of the National Academy of Sciences of the United States of America 114 (27): 7061–7066. DOI: 10.1073/pnas.1616755114 - …

Dr. Simon Aeschbacher
University of Zurich
Department of Evolutionary Biology and Environmental Studies
8057 Zurich
Tel: +41 (0)44 635 49 72
Interdisciplinarity
- Mathematics and biology
- Theoretical and applied population genetics